 I need to dig into CodeRunner a bit more deeply, I think, to get a feel for what gets passed where, to see how a Jupyter sandbox might fit. If a test cell raises an exception, the student gets no marks (or a partial mark in a recent PR) for that question. The notebook is stripped of test/answer cells, provided to the student, completed by them, returned, and then has the test cells added back in. The way that nbgrader (the Jupyter autograder) works is essentially to define a marking notebook that has question cells and a test/answer cell associated with each one. 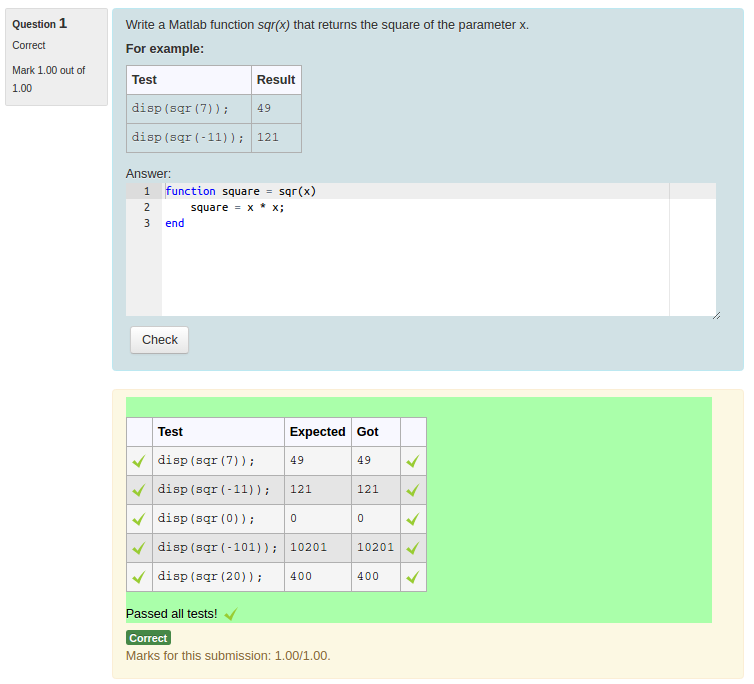
If I wanted to execute a single chunk of code in Jupyter, I could launch a kernel, execute that code chunk there, get the response, and kill the kernel. State is maintained within the environment as long as the kernel process exists.īut that doesn't stop me running a single block of code. (So yes, I can run individual code cells in a notebook UI, in any order, persisting state between code cell executions, and return a response from each code cell execution). 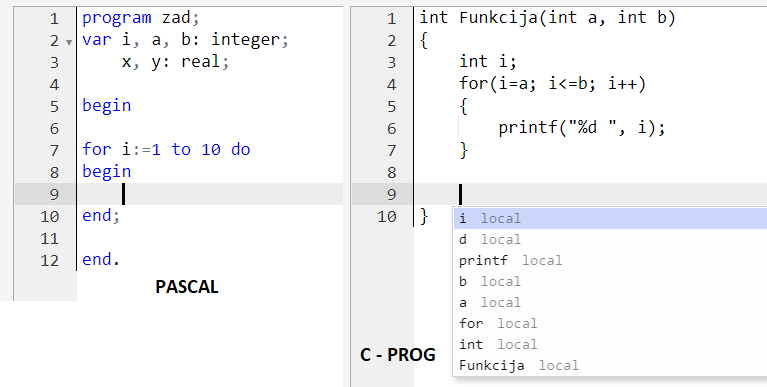
A Jupyter server launches an environment within which you can execute code, returning the any response from the last line of code in the code chunk executed. I don't quite see how a notebook style of programming, which (as I understand it) seems to place a heavy dependence on global state as multiple cells are run, would fit with the CodeRunner model of submitting a complete working standalone program (possibly with support The Jupyter ecosystem is not just about notebooks. I'm not sure that the two would play well together, however. I take your point that if you're teaching with Jupyter and wanted to assess with CodeRunner it would be comforting to know that jobs were being run with exactly the same language software in both cases. Because I don't use Jupyter myself it wasn't a priority. I did look into this briefly but the Jupyter protocol is much more complicated than the Jobe protocol (which is documented here). It would certainly be feasible to use a Jupyter server as a sandbox for CodeRunner by writing another sandbox plugin. By default the first available sandbox that supports the language required by the question is used. CodeRunner can support multiple sandboxes at the same time and questions can be configured to select a particular sandbox (if desired). The latter interfaces to the Sphere On-line judge server but is now more-or-less defunct. Several different ones have been used over the years but the only current ones are the jobe sandbox (file jobesandbox.php) and the ideone sandbox. 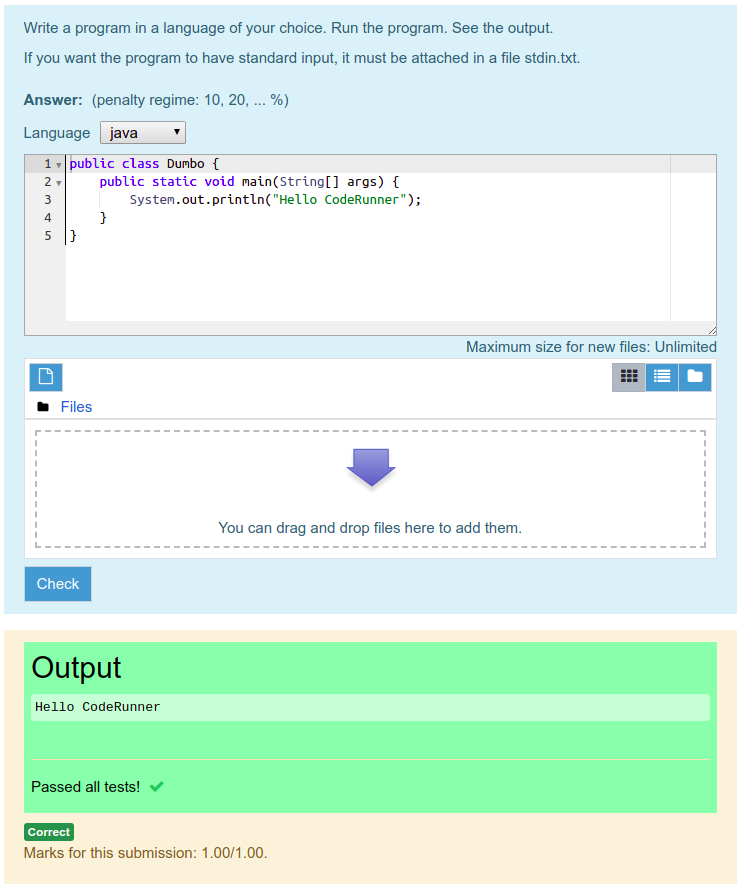
Sandboxes are essentially plugins to CodeRunner. Internally CodeRunner is designed to support multiple sandboxes, implemented as subclasses of the abstract class qtype_coderunner_sandbox - see sandbox.php. (As an aside, Jupyter has several candidate autograding solutions, such as nbgrader as well as integration with okpy, but nothing that I am aware of that is integrated with Moodle explicitly.) This would allow Jupyter users to make use of the same code environments for running coding labs using Jupyter UIs as well as CodeRunner based assessments, as well as extending the range of code environments supported by CodeRunner. If the coupling is handled via a well defined API, then could a middleware shim be used to provide sandboxed code execution either via remotely launched Jupyter kernels or via the current sandboxed environment server? Which got me wondering: how well defined is the API to the sandbox environment which CodeRunner executes code in and is the sandbox server ( Jobe?) expected to run as service? (Indeed, can it be run as a remote service?) One of the issues with using different code environments for teaching (eg using Jupyter notebooks) and assessment (eg using CodeRunner) is the need to maintain consistency across code environments if students are expected to create assessed code fragments that run the same way in the test server as their own Jupyter powered coding environments. Jupyter servers such as the Jupyter kernel gateway or the Jupyter enterprise gateway provide headless access to individually launched remote kernels. Although most commonly known as an interactive notebook serving environment, the Jupyter ecosystem may be more usefully thought of as a set of protocols that, among other things, support remote code execution in dynamically launched "kernels" which are now available for a wide variety of programming languages. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
